 If required, index files can be built from a reference sequence (in FASTA format) using the following command: scratch/work/cgsb/gencore/data/variant_calling/ref/prebuilt/ Reference index files for the sample data have been prebuilt and are available in: Note: Most aligners require an indexed reference sequence as input. 75bp and up.Īlternative aligners such as Bowtie2 may be used. Note that BWA MEM is recommended for longer reads, ie. We use BWA MEM because it is recommended in the Broads best practices and because it has been found to produce better results for variant calling. 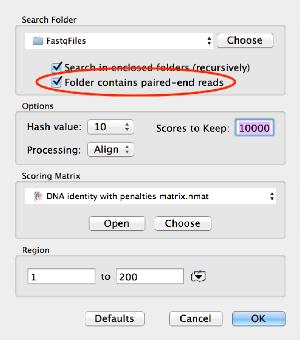
We will use the BWA MEM algorithm to align input reads to your reference genome. 
Prepare reference dictionary, fasta index, and bam indexġ) The Burroughs Wheeler Transform 2) Performing a read alignment using Illumina data.Sort sam file (output from alignment) and convert to bam.This module describes how to map short DNA sequence reads, assess the quality of the alignment and prepare to visualize the mapping of the reads. Once data are in a FASTQ format the first step of any NGS analysis is to align the short reads against the reference genome. JBrowse: Visualizing Data Quickly & Easily.Loading your own data in Seurat & Reanalyze a different dataset.Seurat part 3 – Data normalization and PCA.Exercise part4 – Alternative approach in R to plot and visualize the data.Deeptools2 computeMatrix and plotHeatmap using BioSAILs.Prerequisites, data summary and availability.Instructions to install R Modules on Dalma.Salmon & kallisto: Rapid Transcript Quantification for RNA-Seq Data.Over-Representation Analysis with ClusterProfiler.Gene Set Enrichment Analysis with ClusterProfiler.NGS Sequencing Technology and File Formats.Next-Generation Sequencing Analysis Resources.A common request, especially in our recent survey, is to align existing primer sequences against a template sequence. There are many ways to do this in MacVector, depending on what your requirements are Using the Find dialogįor quickly finding a single primer in a sequence the Find dialog is the first point of call. This allows you to find any sequence, whether it binds to the complementary strand, it is reversed or both. It also allows you to state which end of the sequence to start from and in which direction to scan the sequence (not necessarily obvious!). The Find dialog (see below) is a little daunting to look at first (in fact due to recent user feedback we will hiding most of the functionality in the next release). (i) open up your sequence and use the menu option EDIT > FIND or use the key combination CMD – F However, it is very powerful and in most cases you do not need to change anything as the default settings will work for the majority of cases. When to use: When you need to find a single primer very quickly and do not need to store the results (ii) Enter your primer sequence in the FIND box and click FIND. (iv) Change the drop down menu to Sequence Confirmation then change these parameters: (i) Open the template file and go to ANALYZE > ALIGN TO REFERENCE If you want to quickly align a large set of primers against a template sequence, then as long as each primer is in a separate MacVector file, or a multi sequence fasta file then you can use Sequence Confirmation in the Align to Reference function: Limitations: You can only find a single primer. – If you suspect your primers may not be a perfect match then reduce SCORE THRESHOLD until your primer aligns. The resulting alignment will show your primers aligned against the template. You can switch to the Map view to show a graphical overview of all your primers and where they are located on the sequence. When to use: When you need to find many primers over a large sequenceīenefits: Easy to visualize a great quantity of primers against a template You can use the Editor view for a sequence level representation of the primer aligned against the template. Limitations: Your primer sequences need to be in a file already. This function is fairly easy to use and gives a large amount of detail about your primers. (i) Open your sequence and go to PRIMERS > Test PCR Primer Pairs For example secondary structure, what size the product will be and even the most ideal Tm for your PCR run. (ii) Copy and paste your two sequences in the two boxes. 
Note you will need to have pasted them into an external application. (iii) Click APPLY and see your primers detailed statistics. Primer 2: score 20, mismatches 0, lower strand 1592 to 1573 The 3' end of the primer binds within the product primer 1: score 20, mismatches 0, upper strand 1055 to 1074 Major product size: 538 bp Product details: (iv) Click OK to see the full statistics on the primers and product. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
